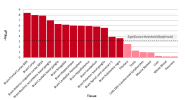
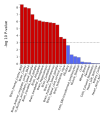
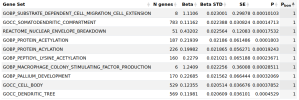
As mentioned above, something
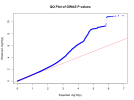
@forestglip and I have been working on recently is running MAGMA locally on the DecodeME summary statistics.
I wouldn’t necessarily recommend anyone do this or follow these instructions. There may be things wrong with this method and it seems to end up with only 94% of variants mapped while
@forestglip gets over 99% using the LiftOver and FUMA method mentioned earlier.
But it was a useful exercise to learn a bit about the process and in sharing this we hope others can learn or indeed point out errors, which helps us learn!
A recipe for gene enrichment with DecodeMe and MAGMA
You will need the following ingredients
- a computer with a shell (performed on Linux but can also be done on Mac or Windows)
- an understanding of installing and running tools from command-line
enough space to download lots of data (15GB or so)
Prepare your tools
- As well as standard tools like awk and zcat you will need to install sort-bed, bedmap and magma itself
Prepare your data
- UCSC genome annotation database for GRCh38/hg38) assembly
We recommend organising these into tools and data folders for easy management, especially if you’ll be running multiple passes with different databases or options. Most need unzipping although the snp151 database is very large so best left zipped.
Then perform the following steps
Filter the main gwas file to include just those variants which passed quality control
Code:
awk -v OFS='\t' 'FNR==NR {ids[$1]++; next} NR==1 || ($3 in ids)' gwas_qced.var gwas_1.regenie > filtered_regenie_file.txt
Add rsid information to these summary stats
First we create then sort a bed formatted file of chromosomes and positions (zero indexed) from the above
Code:
awk -v OFS='\t' 'NR!=1 { print "chr"$1, $2-1, $2, $0 }' filtered_regenie_file.txt > query.bed
sort-bed query.bed > sorted_query.bed
And filter the UCSC genome annotation database to include only single nucleotide variations and again output this in bed format
Code:
zcat snp151.txt.gz | awk -v OFS='\t' '{if($12=="single") print $2,$3,$4,$5}' > snp151_snp.bed
Then we use bedmap to matching rsids to the positions in the summary stats bed file
Code:
bedmap --echo --echo-map-id --delim '\t' sorted_query.bed snp151_snp.bed > sorted_query_with_rsids.bed
Now we prepare the data for MAGMA
First create a gene position file from the above using the rsids as an index
Code:
awk -v OFS='\t' '$21 != "" {print $21,$4,$5}' sorted_query_with_rsids.bed > decodeme.magma_input.id.chr.genpos.txt
Then a p value file
Code:
awk -v OFS='\t' '$21 != "" {print $21,10^(-$19)}' sorted_query_with_rsids.bed > decodeme.magma_input.id.p.txt
Now we can use MAGMA to annotate
Code:
snploc=./decodeme.magma_input.id.chr.genpos.txt
ncbi38=./NCBI38.gene.loc
magma --annotate \
--snp-loc ${snploc} \
--gene-loc ${ncbi38} \
--out decodeme
Then perform gene based analysis
Code:
ref=./g1000_eur/g1000_eur
magma \
--bfile $ref \
--pval ./decodeme.magma_input.id.p.txt N=275488 \
--gene-annot decodeme.genes.annot \
--out decodeme
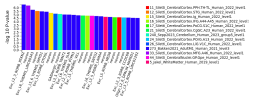
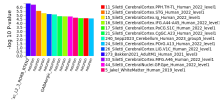
And finally gene-set analysis
Code:
geneset=msigdb_v2025.1.Hs_GMTs/c5.all.v2025.1.Hs.entrez.gmt
magma \
--gene-results decodeme.genes.raw \
--set-annot ${geneset} \
--out decodeme
After a short wait you will have some delicious data to consume with friends!
Thanks to those who made and documented these tools
https://cncr.nl/research/magma/
https://bedops.readthedocs.io/en/latest/
As well the wise and informative internet for its advice on how to use them
https://cloufield.github.io/GWASTutorial/09_Gene_based_analysis/
https://hanbin973.github.io/bioinformatics/2023/02/20/rsid.html
There’s also a handy bash script attached, which will ask for paths to the relevant locations and run it all for you. This also saves configurations and allows reuse of some intermediate files so makes multiple runs with slightly different settings or data easier. The script was put together with help from Gemini Pro for all the boring scaffolding.