Covidivici
Established Member (Voting Rights)
Microbiota-induced intestinal barrier disruption drives BAFF-mediated B-cell dysregulation and autoimmunity in long COVID
Abstract
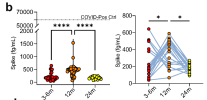
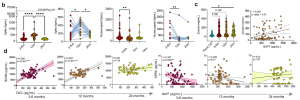
Abstract Long COVID (post-coronavirus disease 2019 (COVID-19) condition) is an infection-associated chronic condition that can follow severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection and persist for months to years1-3. Symptoms can be severe and significantly impact functional status and quality of life2,4,5, but the mechanisms linking barrier dysfunction to systemic immune dysregulation remain unclear. Markers consistent with impaired intestinal mucosal barrier integrity and microbial translocation have been linked to long COVID6. Here we show that non-hospitalized individuals with long COVID have intestinal barrier dysfunction associated with increased B-cell activating factor (BAFF), perturbation of the B cell compartment and autoimmunity that peak at 12 months after infection and begin to resolve by 24 months. Transfer of faecal microbiota from individuals with severe long COVID into germ-free mice is sufficient to reproduce intestinal barrier dysfunction, systemic immune dysregulation with autoimmunity, and neuroinflammation. Treating recipient gnotobiotic mice with a BAFF-neutralizing monoclonal antibody, an approach supported by BAFF biology and clinical efficacy in autoantibody-mediated disease7,8, markedly improves these abnormalities. Together, these findings implicate microbiota-linked intestinal barrier disruption as a driver of autoimmunity and end-organ complications in long COVID and identify BAFF as a therapeutic target.
Web | DOI | Research Square | Preprint (Under Review by Nature Portfolio)
Doyon-Laliberté, Kim; Aranguren, Matheus; Gao, Fandi; Leclerc, Laurence; Fourcade, Lyvia; Patel, Suhani; DuSablon, Charlotte; Conde, Estefania Rivera; Munoz, Diana Cabrera; Purchase, Ludhovik; Nayyerabadi, Maryam; Villard, Alexandre; Darbinian, Emma; Desjardins, Aléhandra; Martineau, Evelyne; Paul, Marie-Lorna; Joshi, Swarali; Mlaga, Kodjovi; Massé, Chantal; Chandrasekaran, Prabha; Poudrier, Johanne; Falcone, Emilia
Abstract
Abstract Long COVID (post-coronavirus disease 2019 (COVID-19) condition) is an infection-associated chronic condition that can follow severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection and persist for months to years1-3. Symptoms can be severe and significantly impact functional status and quality of life2,4,5, but the mechanisms linking barrier dysfunction to systemic immune dysregulation remain unclear. Markers consistent with impaired intestinal mucosal barrier integrity and microbial translocation have been linked to long COVID6. Here we show that non-hospitalized individuals with long COVID have intestinal barrier dysfunction associated with increased B-cell activating factor (BAFF), perturbation of the B cell compartment and autoimmunity that peak at 12 months after infection and begin to resolve by 24 months. Transfer of faecal microbiota from individuals with severe long COVID into germ-free mice is sufficient to reproduce intestinal barrier dysfunction, systemic immune dysregulation with autoimmunity, and neuroinflammation. Treating recipient gnotobiotic mice with a BAFF-neutralizing monoclonal antibody, an approach supported by BAFF biology and clinical efficacy in autoantibody-mediated disease7,8, markedly improves these abnormalities. Together, these findings implicate microbiota-linked intestinal barrier disruption as a driver of autoimmunity and end-organ complications in long COVID and identify BAFF as a therapeutic target.
Web | DOI | Research Square | Preprint (Under Review by Nature Portfolio)