hotblack
Senior Member (Voting Rights)
That’s very positive news! Thanks for doing this @Jonathan Edwards
And good probing questions @voner
And good probing questions @voner
what did you learn that can be shared with us?
Which is really good news!That some other people think this is really interesting!
Or is it more of a pointer to the general pathology
SourceEnhancers and promoters are gene-regulatory elements. They are stretches of DNA that help in both eukaryotic and prokaryotic transcription. The promoters are known to initiate transcription, and the enhancers increase the level of transcription
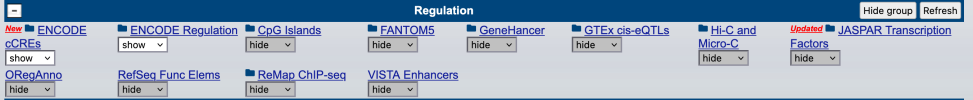
Thanks! The highlighting doesn’t seem very touchscreen friendly unfortunately. But the rest seems to work, I’ll have an explore.@hotblack you might like to explore some of the tracks on UCSC genome browser--it compiles a lot of this information visually so you can do some exploring.
For instance pick the top one, a promoter/enhancer with a high score (it has a little star by it to show this too)
Expand the info and see the location of it is chr17:52158438-52159092
You can zoom in to that location on the DecodeME LocusZoom data and see lots of the hits tie in with this area
Thanks for the tip!If you start zoomed in on a SNP, you can also hit the "highlight" button down here which will highlight everything in your viewer
Good to know I’m not talking nonsense! And nice to start to understand it a bit more.This is more or less exactly what my genetics friends at UCL did when they gave me a presentation of why they thought it was worth picking CA10 for a basic biology project.
Ensembl pipelines are very complex and it would be impossible to provide explanations for all changes between the releases. Sometimes it's possible to figure out or speculate the reasons by looking into intermediate outputs that people who ran the pipelines have but even that is quite challenging. The sheer amount of data and analyses that go into each release... It takes 3-4 or more months and something like 80-100 people working full-time to get a release out. The documentation on the website could be better, though, but it still wouldn't be enough to determine the exact reason behind each change.Wasn't kept in GRCh38, yes--unfortunately Ensembl doesn't really have detailed annotation for why certain genes get dropped in the newest release. Sometimes it's because the gene mapping is suspect, sometimes it's for some other logistical reason. I think that happens a lot to snoRNAs and miRNAs in particular just because of the sheer number of them. But that was the reasoning for creating the Archive--the current version is curated with the best intentions, but shouldn't be considered the end all be all.
The variants are in the current Ensembl release: rs11079993, rs967823. When I clicked on "Phenotype data", an additional piece of info (after a table of associations) said "This variant has not been mapped to any Ensembl genes." for both of them.[
{
"Loci ": "CA10; snoZ178",
"Publication Loci ": "CA10; snoZ178",
"Variants ": "rs11079993",
"allele 1 ": "G",
"allele 2 ": "T",
"direction ": "down",
"Phenotype ": "Other Clinical Pain",
"PMID ": "PMID:33830993",
"comments ": "Significantly associated with multisite chronic pain in female"
},
{
"Loci ": "snoZ178",
"Publication Loci ": "snoZ178",
"Variants ": "rs967823",
"allele 1 ": "G",
"allele 2 ": "A",
"direction ": "no direction reported",
"Phenotype ": "Pain",
"PMID ": "PMID:37844115",
"comments ": "Significntly ssocited with pain"
}
]
library(coloc)
library(dplyr)
library(data.table)
region <- list(chr = 17, start_pos = 52147538, end_pos = 52264626) # CA10 locus
decodeme_region <- fread("../../Data/gwas_1.filtered.gz",
select = c("CHROM", "GENPOS", "ID", "ALLELE0", "ALLELE1",
"A1FREQ", "BETA", "SE", "LOG10P")
) %>%
filter(CHROM == region$chr, GENPOS >= region$start_pos, GENPOS <= region$end_pos) %>%
mutate(
MAF = pmin(A1FREQ, 1 - A1FREQ)
)
gettingup_region <- fread(
"~/Projects/science/diseases/sleep/Jansen 2019/Ease of getting up/Data/Jansen_2019_Gettingup_GRCh38_liftover.tsv.gz",
select = c("SNP", "CHR", "BP", "A1", "A2",
"MAF", "OR", "SE", "P")
) %>%
filter(CHR == region$chr, BP >= region$start_pos, BP <= region$end_pos) %>%
rename(BETA = OR) # OR column appears to actually be BETA, as some values are negative
merged <- inner_join(
decodeme_region %>%
select(GENPOS, ALLELE0, ALLELE1, BETA, SE, MAF),
gettingup_region %>%
select(SNP, BP, A1, A2, BETA, SE, MAF),
by = join_by(GENPOS == BP),
suffix = c("_decodeme", "_gettingup")
) %>%
mutate(
alleles_match = (ALLELE0 == A2 &
ALLELE1 == A1),
alleles_flipped = (ALLELE0 == A1 &
ALLELE1 == A2),
alleles_ok = alleles_match | alleles_flipped,
BETA_gettingup = if_else(alleles_flipped,
-BETA_gettingup,
BETA_gettingup),
) %>%
filter(alleles_ok) %>%
select(SNP, GENPOS,
BETA_decodeme,
BETA_gettingup,
MAF_decodeme,
MAF_gettingup,
SE_decodeme,
SE_gettingup,
)
dataset1 <- list(
beta = merged$BETA_decodeme,
varbeta = merged$SE_decodeme^2,
snp = merged$SNP,
position = merged$GENPOS,
type = "cc"
)
dataset2 <- list(
beta = merged$BETA_gettingup,
varbeta = merged$SE_gettingup^2,
snp = merged$SNP,
position = merged$GENPOS,
type = "quant",
N = 384689,
MAF = merged$MAF_gettingup
)
check_dataset(dataset1)
check_dataset(dataset2)
plot_dataset(dataset1, main = "DecodeME")
plot_dataset(dataset2, main = "Ease of Getting up")
plot_datasets(dataset1, dataset2)
coloc_result <- coloc.abf(
dataset1 = dataset1,
dataset2 = dataset2
)
print(format(round(coloc_result$summary, 3), scientific = FALSE))
head(coloc_result$results[order(coloc_result$results$SNP.PP.H4, decreasing = TRUE), ])I haven't tried downloading it to compare directly, but here is the paper's plot of the locus:There seems to be another more recent study of 'coathanger pain' again using the Biobank that came up with CA10:

A genome-wide association study identifies novel genetic variants associated with neck or shoulder pain in the UK biobank (N = 430,193) - PubMed
In summary, this study has identified novel genetic variants for neck or shoulder pain. Sex-stratified GWAS also suggested that sex played a role in the occurrence of the phenotype.pubmed.ncbi.nlm.nih.gov
Yiwen Tao , Qi Pan, Tengda Cai et al. A genome-wide association study identifies novel genetic variants associated with neck or shoulder pain in the UK biobank (N = 430,193)
Pain Rep 2025 Apr 18;10(3):e1267.
doi: 10.1097/PR9.0000000000001267. eCollection 2025 Jun.
So I'm not sure, but maybe the variant is shared with this trait as well.
Study Trait P-value Sample size 30804565 Ease of getting up in the morning 2.01E-09 385949 31427789 Getting up in morning 3.47E-09 385494 31427789 Overall health rating 9.40E-09 384850 BioRxiv: https://doi.org/10.1101/261081 Ever smoker 3.14E-08 518633 29844566 Reaction time 6.97E-08 330069 31427789 Time spent watching television (TV) 1.86E-07 365236 30696823 Chronotype 5.90E-07 449732 27864402 Self-rated health 8.16E-07 111749 31427789 Pain type(s) experienced in last month: Neck or shoulder pain 0.000001609 385698 31427789 Reaction time test - Mean time to correctly identify matches 0.000001731 383748 31427789 Number of self-reported non-cancer illnesses 0.000001853 386581 30804565 Morningness 0.000002321 345552 31427789 Medication for pain relief, constipation, heartburn: Paracetamol 0.00000241 382089 31427789 Number of treatments/medications taken 0.000002792 386581 31427789 Morning/evening person (chronotype) 0.000002917 345148 31427789 Frequency of tiredness / lethargy in last 2 weeks 0.000002977 375053 30643251 Ever smoked regulary 0.00000572 262990 29899525 Strenuous sports or other exercises 0.00001 350492 31427789 Wheeze or whistling in the chest in last year 0.00001305 379150 31427789 Types of physical activity in last 4 weeks: Other exercises (eg: swimming, cycling, keep fit, bowling) 0.00001679 384450
I'm not understanding why the x axis in the neck shoulder pain chart is different. Can you help me understand?haven't tried downloading it to compare directly, but here is the paper's plot of the locus:
