Jonathan Edwards
Senior Member (Voting Rights)
So it seems there is a shared variant associated with ME/CFS, multisite chronic pain, and ease of getting up in the morning.
Fascinating. Keep at it.
So it seems there is a shared variant associated with ME/CFS, multisite chronic pain, and ease of getting up in the morning.
It's using an older assembly/coordinate system called GRCh37. The more recent one that DecodeME uses is called GRCh38. Defining where exactly a SNP is on a chromosome isn't an exact science, so as they learn more, they make updates to the positions.That's an amazing match with the 'Ease of getting up in the morning' gene. Very impressive investigation FG.
I'm not understanding why the x axis in the neck shoulder pain chart is different. Can you help me understand?
Liftover
This variant lifts over to the following GRCh38 variant:
- 17-52181782-A-C
View variant in gnomAD v4.1.1
Thank you for finding/sharing this. These results are so fascinating. I hope I get less foggy soon so I can read more of the details.The trait most significantly associated with this SNP is "Ease of getting up in the morning", which would make sense as being related to ME/CFS.
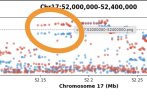
(Maybe this question translates to is 'is this what linkage disequilibrium looks like in summary statistics'?)
Yes, this is showing linkage disequilibrium. The following plot actually shows the strength of LD between each of the variants in the plot with the lead variant (purple diamond).(Maybe this question translates to is 'is this what linkage disequilibrium looks like in summary statistics'?)

Yes, if the causal variants in the two studies were two different high LD "red" variants, the plots would probably look pretty similar. In theory, coloc helps to mathematically determine the probability that there is a shared variant based on the overall pattern, which may subtly change even if the other study's causal variant is a high LD "red" variant.(In that case I guess any condition where one of the dots is elevated you'd expect to see them all elevated?)
id trait chr position (GRCh37) rsid ea nea eaf beta se p n ukb-b-19373 Duration to first press of snap-button in each round 17 50260366 rs34626694 T C 0.328517 0.0151902 0.00217161 2.70023e-12 459281 ukb-b-16287 Mean time to correctly identify matches 17 50260366 rs34626694 T C 0.328492 0.0146137 0.00218135 2.09991e-11 459523 ukb-b-6306 Overall health rating 17 50260366 rs34626694 T C 0.328511 0.0103603 0.00159992 9.3994e-11 460844 ukb-b-2772 Getting up in morning 17 50260366 rs34626694 T C 0.32854 -0.0109886 0.00170231 1.09999e-10 461658 ukb-b-9130 Pain type(s) experienced in last month: None of the above 17 50260366 rs34626694 T C 0.328525 -0.00636032 0.00108326 4.30002e-09 461857 ukb-a-10 Getting up in morning 17 50260366 rs34626694 T C 0.330974 -0.0112884 0.00198473 1.28911e-08 336501 ukb-a-251 Overall health rating 17 50260366 rs34626694 T C 0.330974 0.0102797 0.0018765 4.30229e-08 336020 ukb-b-929 Frequency of tiredness / lethargy in last 2 weeks 17 50260366 rs34626694 T C 0.32834 0.0101412 0.00186099 5.1e-08 449019 ukb-b-18335 Wheeze or whistling in the chest in last year 17 50260366 rs34626694 T C 0.328588 0.00487348 0.000904747 7.19996e-08 453959 ukb-b-8746 Illnesses of siblings: High blood pressure 17 50260366 rs34626694 T C 0.32859 0.00542589 0.00101614 9.29994e-08 364661 ukb-a-199 Mean time to correctly identify matches 17 50260366 rs34626694 T C 0.330974 0.0134514 0.00255376 1.38535e-07 335139 ukb-b-17595 Medication for pain relief, constipation, heartburn: Paracetamol 17 50260366 rs34626694 T C 0.328477 0.00467076 0.000916774 3.50002e-07 457547 ebi-a-GCST90029014 Smoking status 17 50260366 rs34626694 T C 0.328868 0.00734503 0.00146598 3.79997e-07 468170 ukb-d-20116_0 Smoking status: Never 17 50260366 rs34626694 T C 0.331077 -0.00626354 0.00123574 4.00876e-07 359706 ukb-b-18596 Pain type(s) experienced in last month: Neck or shoulder pain 17 50260366 rs34626694 T C 0.328525 0.00470001 0.000933439 4.79999e-07 461857 ebi-a-GCST90012794 Participation in an health questionnaire (not invited vs invited) 17 50260366 rs34626694 T C 0.330099 0.0047584 0.000952877 6.1e-07 451097 ukb-a-472 Pain type(s) experienced in last month: Neck or shoulder pain 17 50260366 rs34626694 T C 0.330974 0.00538691 0.00108634 7.09741e-07 336650 ukb-b-4063 Number of self-reported non-cancer illnesses 17 50260366 rs34626694 T C 0.328506 0.00897475 0.00182012 8.19993e-07 462933
Thanks for doing this.Here are all associations with p<1e-6, sorted starting from most significant:
Yes good point. We can at least see the total sample size for studies in the table above, and they're all around 350,000 to 450,000 for this group of datasets. Though unbalanced group sizes in binary traits, for example, could make a study's effective sample size smaller, making the findings less directly comparable to each other.One issue I see is that we don't have a standardized measure of effect size to filter these. I suspect that in a lot of the things that come up like smoking, overall health and cognitive tests, the DNA has only very minor effects and that they only come up because these traits could be tested with enormous sample sizes. So a large proportion of all possible DNA variants and genes are likely associated with them.

Trait Data type Gene Tissue Cis/Trans P-value ME/CFS 2.11e-9 CA10 Whole blood cg04881814 methQTL Methylation CA10 Whole blood cis 0.00e+0 CA10 Whole blood cg07398767 methQTL Methylation CA10 Whole blood cis 4.19e-14 CA10 Whole blood cg08605326 methQTL Methylation CA10 Whole blood cis 4.10e-19 CA10 Whole blood cg20552747 methQTL Methylation CA10 Whole blood cis 1.87e-15 Gastroesophageal reflux disease Phenotype (Disease Of Digestive System) 1.69e-9 Participation in an health questionnaire (not invited vs invited) Phenotype (Physiological Measures) 1.20e-7 Mean time to correctly identify matches Phenotype (Behavioural Measures) 2.10e-11 Illnesses of siblings: None of the above (group 1) Phenotype (Environmental Measures) 1.20e-7 Medication for pain relief, constipation, heartburn: Paracetamol Phenotype (Physiological Measures) 4.60e-8 Wheeze or whistling in the chest in last year Phenotype (Physiological Measures) 7.20e-8 Pain type(s) experienced in last month: Neck or shoulder pain Phenotype (Disease Of Musculoskeletal System And Connective Tissue) 4.80e-12 Getting up in morning Phenotype (Behavioural Measures) 8.10e-11 Time spent watching television (TV) Phenotype (Behavioural Measures) 5.50e-8 Overall health rating Phenotype (Physiological Measures) 2.60e-12 Illnesses of siblings: High blood pressure Phenotype (Disease Of Circulatory System) 3.00e-8
Info column for this locus says:
Candidate Variant: 17:52176967 A/G
LD Region: EUR/17/49918934-53040813
The formula isNote: Often the effective sample size is defined as 4nϕ(1−ϕ), because that quantity tells what would be the total sample size (cases + controls) in a hypothetical study that has equal number of cases and controls and whose power matches the power of our current study.
Neff = 4 * [total sample size] * [proportion of cases] * [proportion of controls]. For DecodeME, that's Neff = 4 * 275488 * 0.056550558 * 0.943449442 ≈ 58792Neff = 4 / (1/[num cases] + 1/[num controls])Just seeing what's in this paper about CA10.Yiwen Tao , Qi Pan, Tengda Cai et al. A genome-wide association study identifies novel genetic variants associated with neck or shoulder pain in the UK biobank (N = 430,193)
I don't see a "shoulder issues" trait on FinnGen's website, so I'm not sure if it's called something else.In addition, another limitation is particularly notable as we had to use a different phenotype (shoulder issues) instead of neck or shoulder pain. This substitution may have introduced a bias, as neck or shoulder pain, while a prominent indicator, is not entirely synonymous with shoulder issues.
CA10 encodes a protein from the zinc metalloenzyme family called carbonic anhydrase, involved in bone resorption and bone mineral solubilization.38,39
The protein is also believed to be involved in the development of the brain and the central nervous system (https://www.ncbi.nlm.nih.gov/gene/105371829#gene-expression).
The strong interconnections between carbonic anhydrase activity and neuropathic pain have been suggested.3,4 It has been identified that inhibition of carbonic anhydrase activity by intraperitoneal injection of acetazolamide decreased neuropathic pain.7
For CA10, one study suggested that it may play the role of adaptors, facilitating neurexins' indirect connections with unidentified postsynaptic target molecules, and thus mediating the development of new transsynaptic complexes.51
The strong interconnections between carbonic anhydrase activity and neuropathic pain have been suggested.3,4 It has been identified that inhibition of carbonic anhydrase activity by intraperitoneal injection of acetazolamide decreased neuropathic pain.7
Carbonic anhydrases (EC 4.2.1.1) are ubiquitous enzymes that play a key role in acid–base homeostasis by catalyzing the interconversion of carbon dioxide and water to bicarbonate. Carbonic anhydrase-related proteins (CARPs) belong to the family of α-carbonic anhydrases but are catalytically inactive due to the loss of one or more of the histidine residues that coordinate the zinc atom in the catalytic core (15, 16).
34 genes were identified as potentially important in the evolution of the Anxi breed:Based on the comparative genomic selection signal analysis between Anxi cattle and Japanese cattle, this study employed four methods: Fst, θπ, XP-CLR, and XP-EHH for detecting selection signals. [...] Through multi-method cross-validation, 34 high-confidence candidate genes supported by at least three methods were screened out
FBXL17
CA10
CA8
CENPP
CSMD1
CTNNA2
EXOC4
FOXP2
GPC5
HNF4G
LOC100295832
LOC104975749
LOC112444478
LOC112446063
LOC112447533
LOC112447589
LOC112447953
LOC112448128
LOC112448167
LOC112448942
LOC112448971
LOC132342562
LOC132342819
LOC132342820
LOC132343034
LOC132345190
LOC132345454
LOC132346165
LOC786256
PTPRD
SLC16A7
TOX
TRNAS-GGA_96
TRNAY-GUA
Anxi cattle are a superior local population derived from Mongolian cattle. Despite their small body size and relatively slow growth rate, they demonstrate strong adaptability to arid and extreme environmental conditions, making them an essential livestock resource in grassland and desert regions.
Here we used whole genome sequencing data of 104 animals to delve into the population structure, genomic diversity and potential positive selection signals in Zhaotong cattle.
Genome-wide selection scans detected a series of positive candidate regions containing multiple key genes related to bone development and metabolism (CA10, GABRG3, GLDN and NOTUM)
This study compared the genomes of 30 newly sequenced Ningqiang ponies with those of 56 other ponies and 104 horses to investigate genetic diversity, genetic differentiation, and the genetic basis of body height differences.
Additionally, TBX5, ASAP1, CDK12, CA10, and CSMD1 were identified as important candidate genes for body height differences between ponies and horses.
This is the first study that seeks to access such information for the Mangalarga Marchador (MM) horse breed on a genomic scale.
Three regions from the ROHet [runs of heterozygosity] analyses showed genes with known functions: tripartite motif-containing 37 (TRIM37), protein phosphatase, Mg2+/Mn2+ dependent 1E (PPM1E) and carbonic anhydrase 10 (CA10).
regions with high variability in the MM genome have been identified (ROHet), where genes in these regions are possibly associated with recent selection, acting in important events for the development and performance of the MM horse over generations.
CA10 seems to be present in a lot of species. (Here's a gene tree from Ensembl. If it's not expanded, you can click on "View fully expanded tree" under the image.)For some reason, CA10 comes up multiple times in selection analysis of various livestock. Though I have very little understanding of what these results mean. There's the above paper on Anxi cattle, and here are three more on other animals:
...
Edit: There's always the possibility that thousands of these studies on livestock have been done, with all variety of genes found in different studies, and I just picked the few that have CA10.
Baseline ventricular lactate measured by proton magnetic resonance spectroscopic imaging was found to be higher in patients with FM than in healthy controls (F1,37 = 22.11, P < .0001, partial η2 = .37).
There was no significant difference in ventricular lactate levels between patients with FM + CFS compared with those with FM only (n = 17, 8.09 ± 1.85 i.u. [SD] and n = 6, 9.44 ± 1.9 i.u., respectively; F1,19 = 1.03, P = .32).
This might explain the reduction of lactate and glutamate in the DLPFC of chronic back pain subjects (not significant in this study)
Specific activation of dorsal horn astrocytes induced mechanical hyperalgesia, which was attenuated by α-cyano-4-hydroxycinnamate (4-CIN), an inhibitor of monocarboxylate transporters that deliver L-lactate from astrocytes to neurons.
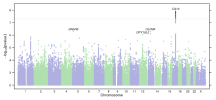

Our GWASes of susceptibility to meningitis (based on a sample of 842 cases) failed to replicate the findings at CFH, but identified a significant association at CA10. Moreover, our case definition of self-reported ‘meningitis’ may be confounded by the fact that bacterial meningitis is frequently considered in patients with presenting to the hospital with severe headache and fever, a definitive diagnosis is made in a minority of cases55. In other words, patients initially suspected to have bacterial meningitis may ultimately be diagnosed with other syndromes such as viral meningitis, stroke, subdural empyema and cerebral abscess55.
Ive just realised I don't know whether CA10 is raised or lowered in DecodeME and the fibro studiesCA10 may play a role in ME/CFS and pain (GWAS).
