I previously did that
colocalization testing using the tool on Open GWAS, and it showed that the ME/CFS CA10 locus colocalizes with a few methQTLs (variants which cause differential methylation at a particular place on the genome):
I think the methQTL data is from the GoDMC project. I searched for the lead ME/CFS CA10 SNP on there, and it showed that it is associated with three of the four methylation sites above:
https://mqtldb.godmc.org.uk/search?query=rs34626694
Here's the data from that search:
So the first of these, cg04881814, is extremely significant compared to the others, with p=7.1e-242. The beta is negative for the T allele (the ME/CFS risk allele). The direction column shows the direction of methylation for each of the cohorts that were tested in this database, and it shows negative for almost all of them. I think this means there is a strong association between the T allele and demethylation at this site.
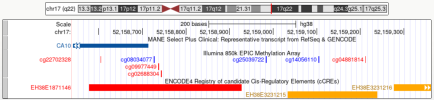
Here is that methylation site, cg04881814, on
UCSC Genome Browser. It's the one furthest to the right, and it overlaps the enhancer EH38E3231215 near the start of CA10.
View attachment 32335
The genome browser links to this website for that enhancer:
https://screen.wenglab.org/GRCh38/ccre/EH38E3231215
I don't know what else to do with this information, or how to interpret what's on that enhancer page.